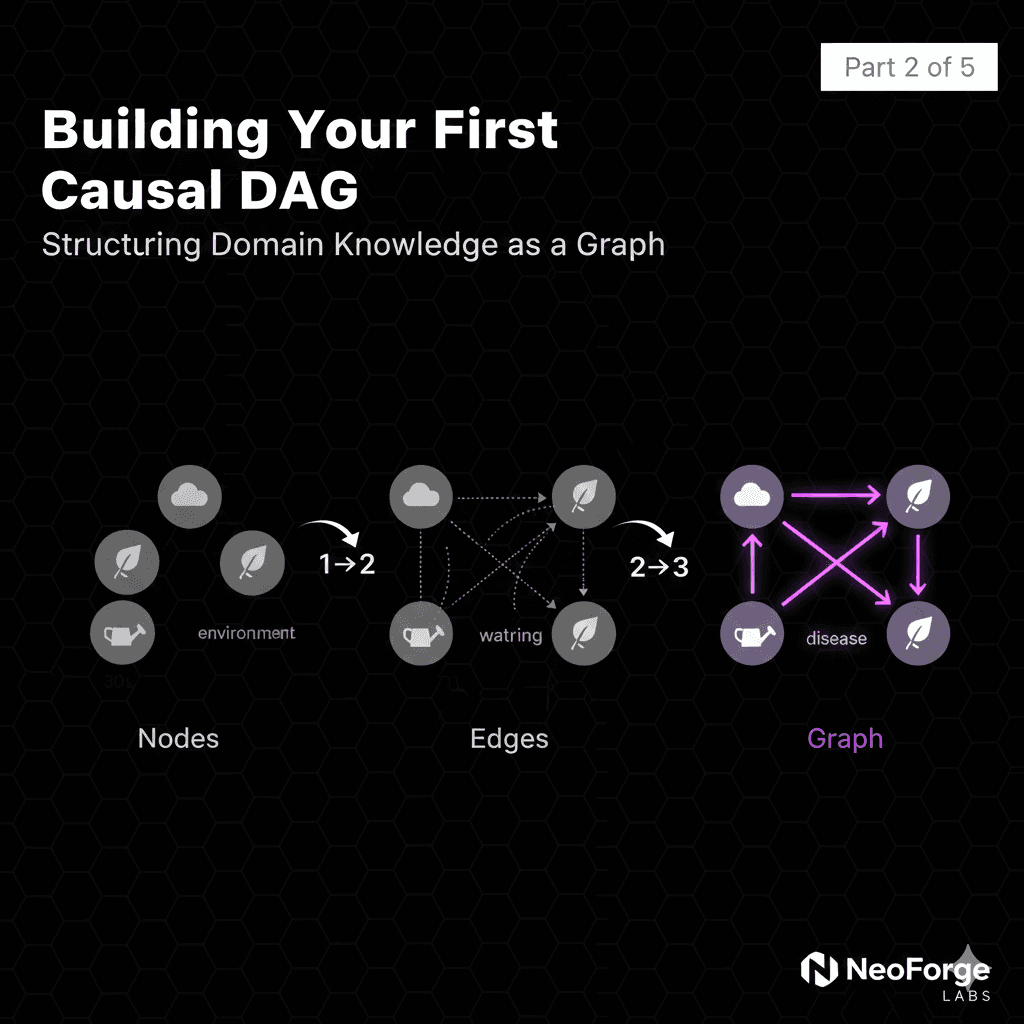
Part 3: Counterfactual Reasoning with Causal DAGs
Predicting Alternate Realities
Senior Software Engineer with a knack for Python, Golang, TypeScript, and Elixir. I am also a bit of a Rust enthusiast. I am excited by all things scalability and microservices. Join me on this journey to becoming a unicorn 10x Engineer.
In Parts 1 and 2, you learned why causality matters and how to build causal DAGs. Today, we're climbing to Level 3 of Pearl's Ladder: Counterfactual Reasoning.
This is the most powerful form of causal AI—reasoning about alternate realities and answering "what if" questions that standard ML can't touch.
By the end of this article, you'll:
Understand what counterfactuals are and why they're powerful
Implement counterfactual inference with your DAG
Generate personalized explanations for individual cases
Estimate individual treatment effects
Build "what if" scenario analysis tools
Let's reason about alternate realities.
What Are Counterfactuals?
The Three Worlds of Causality
Factual World (What happened):
"I watered this plant heavily"
"The plant developed root rot"
This is observation—what we actually saw
Interventional World (What would happen):
"If I water the next plant moderately, what happens?"
"Disease probability drops to 20%"
This is prediction—what we expect in the future
Counterfactual World (What would have happened):
"Would THIS plant be healthy if I had watered it differently?"
"85% probability it would be healthy"
This is retrospection—alternate history for a specific instance
Why Counterfactuals Are Special
Counterfactuals require THREE pieces of information:
1. The causal mechanism (from your DAG)
How do variables causally relate?
Watering → Moisture → Pathogen → Disease
2. The specific instance (observed data)
This plant had: high watering, high moisture, disease
Its vigor: 0.7, environmental stress: 0.4
3. The alternate action (the intervention)
What if watering had been moderate instead?
How would moisture, pathogen, disease differ?
Standard ML only has #2. Intervention (Level 2) has #1 and #3. Only counterfactuals combine all three.
The Counterfactual Formula
Notation: P(Y_x' | X=x, Y=y)
Read as: "Probability of outcome Y under intervention x', given we observed X=x and Y=y in reality"
Example:
P(Healthy | do(Watering=moderate), Watering=heavy, Diseased)
"Would plant be healthy with moderate watering, given we observed heavy watering and disease?"
This is fundamentally different from:
P(Healthy | Watering=moderate) — observational (correlation)
P(Healthy | do(Watering=moderate)) — interventional (average effect)
Counterfactuals condition on BOTH the alternate intervention AND the factual observation.
Implementing Counterfactual Inference
The Three Steps of Counterfactual Analysis
Step 1: Abduction — Infer latent variables from observations Step 2: Action — Modify the model according to the intervention Step 3: Prediction — Compute the counterfactual outcome
Let's implement this with our plant disease DAG:
import numpy as np
import pandas as pd
from dowhy import CausalModel
class CounterfactualEngine:
"""
Counterfactual reasoning engine for plant disease diagnosis.
"""
def __init__(self, causal_dag, data):
self.dag = causal_dag
self.data = data
self.model = self._build_model()
def _build_model(self):
"""Build the causal model from DAG."""
return CausalModel(
data=self.data,
treatment='leaf_moisture_hours',
outcome='symptom_severity',
graph=self.dag,
common_causes=['environmental_stress', 'watering_practice'],
effect_modifiers=['plant_vigor']
)
def abduction(self, sample_idx):
"""
Step 1: Infer latent variables from observed data.
Given what we observed, what are the unobserved factors?
"""
observed = self.data.iloc[sample_idx]
# Infer noise terms (unobserved confounders)
# These capture the instance-specific factors
noise_terms = {
'u_moisture': self._infer_moisture_noise(observed),
'u_pathogen': self._infer_pathogen_noise(observed),
'u_disease': self._infer_disease_noise(observed),
'u_severity': self._infer_severity_noise(observed)
}
return observed, noise_terms
def _infer_moisture_noise(self, obs):
"""Infer moisture noise from observation."""
# Expected moisture given inputs
expected = 5.0 + obs['environmental_stress'] * 10
if obs['watering_practice'] == 0:
expected -= 3
elif obs['watering_practice'] == 2:
expected += 5
# Noise is observed - expected
return obs['leaf_moisture_hours'] - expected
def _infer_pathogen_noise(self, obs):
"""Infer pathogen growth noise."""
expected = (obs['leaf_moisture_hours'] / 24) ** 1.5
return obs['pathogen_growth'] - expected
def _infer_disease_noise(self, obs):
"""Infer disease threshold noise."""
# Binary outcome, return indicator
return obs['disease_present']
def _infer_severity_noise(self, obs):
"""Infer symptom severity noise."""
if obs['disease_present'] == 0:
return 0
expected = obs['disease_present'] * (1 - obs['plant_vigor'] * 0.5)
return obs['symptom_severity'] - expected
def action(self, observed, noise_terms, intervention):
"""
Step 2: Modify model according to intervention.
Set the treatment variable to counterfactual value.
"""
# Create counterfactual data point
cf_data = observed.copy()
# Apply intervention (break incoming edges)
for var, value in intervention.items():
cf_data[var] = value
return cf_data, noise_terms
def prediction(self, cf_data, noise_terms):
"""
Step 3: Compute counterfactual outcome.
Propagate intervention through causal graph.
"""
# Re-compute downstream variables with intervention
# Leaf moisture (intervened, so use counterfactual value)
cf_moisture = cf_data['leaf_moisture_hours']
# Pathogen growth (function of new moisture + same noise)
cf_pathogen = (cf_moisture / 24) ** 1.5 + noise_terms['u_pathogen']
cf_pathogen = np.clip(cf_pathogen, 0, 1)
# Disease (function of new pathogen + same noise threshold)
cf_disease = 1 if cf_pathogen > 0.6 else 0
# Symptom severity (function of new disease + same plant vigor + same noise)
cf_severity = cf_disease * (1 - cf_data['plant_vigor'] * 0.5) + noise_terms['u_severity']
cf_severity = np.clip(cf_severity, 0, 1)
return {
'leaf_moisture_hours': cf_moisture,
'pathogen_growth': cf_pathogen,
'disease_present': cf_disease,
'symptom_severity': cf_severity
}
def counterfactual(self, sample_idx, intervention):
"""
Complete counterfactual analysis.
Args:
sample_idx: Index of observed instance
intervention: Dict of {variable: counterfactual_value}
Returns:
Dict with factual, counterfactual, and effect
"""
# Step 1: Abduction
observed, noise_terms = self.abduction(sample_idx)
# Step 2: Action
cf_data, noise_terms = self.action(observed, noise_terms, intervention)
# Step 3: Prediction
cf_outcome = self.prediction(cf_data, noise_terms)
return {
'factual': {
'leaf_moisture_hours': observed['leaf_moisture_hours'],
'pathogen_growth': observed['pathogen_growth'],
'disease_present': observed['disease_present'],
'symptom_severity': observed['symptom_severity']
},
'counterfactual': cf_outcome,
'individual_effect': {
'disease_change': cf_outcome['disease_present'] - observed['disease_present'],
'severity_change': cf_outcome['symptom_severity'] - observed['symptom_severity']
},
'explanation': self._generate_explanation(observed, cf_outcome)
}
def _generate_explanation(self, factual, counterfactual):
"""Generate natural language explanation of counterfactual."""
explanation = []
# Compare factual vs counterfactual
if factual['disease_present'] == 1 and counterfactual['disease_present'] == 0:
explanation.append(
f"With moderate watering (reducing moisture from {factual['leaf_moisture_hours']:.1f} to "
f"{counterfactual['leaf_moisture_hours']:.1f} hours), this plant would have avoided disease."
)
elif factual['disease_present'] == 0 and counterfactual['disease_present'] == 1:
explanation.append(
f"If watering had been excessive (increasing moisture to "
f"{counterfactual['leaf_moisture_hours']:.1f} hours), this plant would have developed disease."
)
else:
explanation.append(
f"Disease status would remain unchanged, but symptom severity would change from "
f"{factual['symptom_severity']:.2f} to {counterfactual['symptom_severity']:.2f}."
)
# Add mechanism
explanation.append(
f"Mechanism: Moisture affects pathogen growth ({factual['pathogen_growth']:.2f} → "
f"{counterfactual['pathogen_growth']:.2f}), which determines disease presence."
)
return " ".join(explanation)
# Usage Example
if __name__ == "__main__":
# Load data and DAG (from Part 2)
from part2_causal_dag import generate_causal_data, causal_graph
data = generate_causal_data(n_samples=1000)
cf_engine = CounterfactualEngine(causal_graph, data)
# Find a diseased plant
diseased_idx = data[data['disease_present'] == 1].index[0]
print("=" * 60)
print("COUNTERFACTUAL ANALYSIS")
print("=" * 60)
print(f"\nAnalyzing Plant #{diseased_idx}")
print(f"Factual: Watering = {data.loc[diseased_idx, 'watering_practice']}")
print(f" Moisture = {data.loc[diseased_idx, 'leaf_moisture_hours']:.1f} hours")
print(f" Disease = {bool(data.loc[diseased_idx, 'disease_present'])}")
print(f" Severity = {data.loc[diseased_idx, 'symptom_severity']:.2f}")
# Counterfactual: What if watering was optimal?
intervention = {'leaf_moisture_hours': 6.0} # Optimal moisture
result = cf_engine.counterfactual(diseased_idx, intervention)
print(f"\nCounterfactual: Watering = optimal")
print(f" Moisture = {result['counterfactual']['leaf_moisture_hours']:.1f} hours")
print(f" Disease = {bool(result['counterfactual']['disease_present'])}")
print(f" Severity = {result['counterfactual']['symptom_severity']:.2f}")
print(f"\nIndividual Treatment Effect:")
print(f" Disease change: {result['individual_effect']['disease_change']}")
print(f" Severity change: {result['individual_effect']['severity_change']:.2f}")
print(f"\nExplanation:")
print(f" {result['explanation']}")
Output Example
Applications of Counterfactual Reasoning
1. Personalized Recommendations
Standard ML: "Plants with disease X should receive treatment Y" (average effect)
Counterfactual AI: "THIS plant would benefit most from intervention Z" (personalized)
def recommend_intervention(cf_engine, plant_idx):
"""
Find optimal intervention for specific plant.
"""
# Test multiple interventions
interventions = {
'reduce_watering': {'leaf_moisture_hours': 5.0},
'moderate_watering': {'leaf_moisture_hours': 8.0},
'increase_watering': {'leaf_moisture_hours': 12.0}
}
results = {}
for name, intervention in interventions.items():
result = cf_engine.counterfactual(plant_idx, intervention)
results[name] = result['counterfactual']['symptom_severity']
# Find best intervention
best = min(results.items(), key=lambda x: x[1])
return {
'recommendation': best[0],
'expected_severity': best[1],
'all_options': results
}
# Example usage
plant_idx = 42
recommendation = recommend_intervention(cf_engine, plant_idx)
print(f"Optimal intervention for Plant #{plant_idx}:")
print(f" {recommendation['recommendation']}")
print(f" Expected severity: {recommendation['expected_severity']:.2f}")
print(f"\nAll options:")
for intervention, severity in recommendation['all_options'].items():
print(f" {intervention}: {severity:.2f}")
2. Explanation Generation
Why did this plant get diseased?
def explain_disease(cf_engine, diseased_idx, healthy_idx):
"""
Explain why one plant got diseased and another didn't.
"""
diseased = cf_engine.data.iloc[diseased_idx]
healthy = cf_engine.data.iloc[healthy_idx]
# Compare key differences
differences = []
if diseased['watering_practice'] != healthy['watering_practice']:
differences.append(
f"Watering: Plant #{diseased_idx} was watered differently "
f"({diseased['watering_practice']} vs {healthy['watering_practice']})"
)
if abs(diseased['plant_vigor'] - healthy['plant_vigor']) > 0.2:
differences.append(
f"Vigor: Plant #{diseased_idx} had {'lower' if diseased['plant_vigor'] < healthy['plant_vigor'] else 'higher'} vigor "
f"({diseased['plant_vigor']:.2f} vs {healthy['plant_vigor']:.2f})"
)
# Counterfactual: Would diseased plant be healthy with healthy plant's watering?
intervention = {'leaf_moisture_hours': healthy['leaf_moisture_hours']}
cf_result = cf_engine.counterfactual(diseased_idx, intervention)
if cf_result['counterfactual']['disease_present'] == 0:
differences.append(
f"CRITICAL: If Plant #{diseased_idx} had received the same watering as "
f"Plant #{healthy_idx}, it would have remained healthy."
)
return {
'differences': differences,
'counterfactual': cf_result,
'root_cause': 'watering_practice' if cf_result['individual_effect']['disease_change'] < 0 else 'plant_vigor'
}
# Usage
diseased_plant = data[data['disease_present'] == 1].index[0]
healthy_plant = data[data['disease_present'] == 0].index[0]
explanation = explain_disease(cf_engine, diseased_plant, healthy_plant)
print(f"Why did Plant #{diseased_plant} get diseased?")
for diff in explanation['differences']:
print(f" • {diff}")
print(f"\nRoot cause: {explanation['root_cause']}")
3. Regret Analysis
What should I have done differently?
def regret_analysis(cf_engine, sample_idx):
"""
Analyze what optimal action would have been.
"""
actual = cf_engine.data.iloc[sample_idx]
# Test all possible watering practices
watering_options = [0, 1, 2] # under, optimal, over
results = {}
for watering in watering_options:
# Compute expected moisture for this watering
expected_moisture = 5.0 + actual['environmental_stress'] * 10
if watering == 0:
expected_moisture -= 3
elif watering == 2:
expected_moisture += 5
intervention = {'leaf_moisture_hours': max(0, min(24, expected_moisture))}
cf_result = cf_engine.counterfactual(sample_idx, intervention)
results[watering] = {
'disease': cf_result['counterfactual']['disease_present'],
'severity': cf_result['counterfactual']['symptom_severity']
}
# Find optimal action
optimal = min(results.items(), key=lambda x: (x[1]['disease'], x[1]['severity']))
actual_watering = actual['watering_practice']
regret = {
'optimal_action': optimal[0],
'actual_action': actual_watering,
'regret': results[actual_watering]['severity'] - optimal[1]['severity']
}
return regret
# Usage
plant_idx = 42
regret = regret_analysis(cf_engine, plant_idx)
print(f"Regret Analysis for Plant #{plant_idx}:")
print(f" Actual action: {['under', 'optimal', 'over'][regret['actual_action']]} watering")
print(f" Optimal action: {['under', 'optimal', 'over'][regret['optimal_action']]} watering")
print(f" Regret: {regret['regret']:.2f} severity points")
if regret['regret'] > 0.1:
print(f" ⚠️ Significant regret! Better watering would have reduced severity substantially.")
else:
print(f" ✓ Action was near-optimal.")
4. Policy Evaluation
Was our intervention strategy effective?
def evaluate_policy(cf_engine, treated_indices, control_indices):
"""
Evaluate treatment effect using counterfactual reasoning.
"""
# For treated group: What if they hadn't been treated?
treated_effects = []
for idx in treated_indices:
# Assume treatment was reducing moisture
cf_result = cf_engine.counterfactual(
idx,
{'leaf_moisture_hours': cf_engine.data.loc[idx, 'leaf_moisture_hours'] + 5.0}
)
treated_effects.append(cf_result['individual_effect']['severity_change'])
# For control group: What if they had been treated?
control_effects = []
for idx in control_indices:
cf_result = cf_engine.counterfactual(
idx,
{'leaf_moisture_hours': max(0, cf_engine.data.loc[idx, 'leaf_moisture_hours'] - 5.0)}
)
control_effects.append(-cf_result['individual_effect']['severity_change'])
# Overall treatment effect
ate = np.mean(treated_effects + control_effects)
return {
'average_treatment_effect': ate,
'treated_effect': np.mean(treated_effects),
'control_effect': np.mean(control_effects),
'heterogeneity': np.std(treated_effects + control_effects)
}
Individual Treatment Effects (ITE)
Beyond Average Treatment Effects
Average Treatment Effect (ATE): What's the effect on average?
- "Reducing watering decreases disease by 15% on average"
Individual Treatment Effect (ITE): What's the effect for THIS individual?
"For Plant #42, reducing watering would decrease disease by 85%"
"For Plant #17, reducing watering would have no effect"
Why ITE Matters
Precision medicine/agriculture:
Not everyone responds the same way
Treatment X might help person A but harm person B
Counterfactuals let us estimate personalized effects
Computing ITE
def compute_ite(cf_engine, sample_idx, treatment_var, treatment_value):
"""
Compute Individual Treatment Effect.
ITE = Y_1 - Y_0
where Y_1 is outcome under treatment, Y_0 is outcome under control
"""
# Factual outcome (what actually happened)
factual = cf_engine.data.iloc[sample_idx]
# Counterfactual outcome (what would happen under treatment)
intervention = {treatment_var: treatment_value}
cf_result = cf_engine.counterfactual(sample_idx, intervention)
ite = cf_result['counterfactual']['symptom_severity'] - factual['symptom_severity']
return {
'ite': ite,
'factual_outcome': factual['symptom_severity'],
'counterfactual_outcome': cf_result['counterfactual']['symptom_severity'],
'would_benefit': ite < -0.1, # At least 10% improvement
'confidence': 'high' if abs(cf_result['counterfactual']['pathogen_growth'] - factual['pathogen_growth']) > 0.2 else 'low'
}
# Usage: Estimate ITE for multiple plants
ite_results = []
for idx in range(100):
ite = compute_ite(cf_engine, idx, 'leaf_moisture_hours', 6.0)
ite_results.append({
'plant_idx': idx,
'ite': ite['ite'],
'would_benefit': ite['would_benefit']
})
ite_df = pd.DataFrame(ite_results)
print("Individual Treatment Effect Distribution:")
print(f" Mean ITE: {ite_df['ite'].mean():.3f}")
print(f" Std ITE: {ite_df['ite'].std():.3f}")
print(f" % who would benefit: {ite_df['would_benefit'].mean():.1%}")
# Identify who benefits most
top_beneficiaries = ite_df.nsmallest(10, 'ite')
print(f"\nTop 10 beneficiaries from treatment:")
print(top_beneficiaries)
Counterfactual Fairness
Ensuring Fair AI Decisions
The problem: ML models can discriminate based on protected attributes
Counterfactual fairness: "Would the decision be the same if the person had a different protected attribute?"
def check_counterfactual_fairness(cf_engine, sample_idx, protected_attr, alt_value):
"""
Check if decision would change with different protected attribute.
"""
# Factual decision
factual = cf_engine.data.iloc[sample_idx]
factual_decision = "treat" if factual['symptom_severity'] > 0.5 else "monitor"
# Counterfactual decision (with different protected attribute)
intervention = {protected_attr: alt_value}
cf_result = cf_engine.counterfactual(sample_idx, intervention)
cf_decision = "treat" if cf_result['counterfactual']['symptom_severity'] > 0.5 else "monitor"
is_fair = factual_decision == cf_decision
return {
'is_fair': is_fair,
'factual_decision': factual_decision,
'counterfactual_decision': cf_decision,
'protected_attr': protected_attr,
'explanation': f"Decision {'would' if is_fair else 'would NOT'} remain the same"
}
Practical Tips for Counterfactual Reasoning
1. Validate Structural Equations
Your counterfactuals are only as good as your causal model:
Test on known interventions
Compare to randomized trials when available
Check if counterfactual predictions match observed data
2. Handle Uncertainty
def counterfactual_with_uncertainty(cf_engine, sample_idx, intervention, n_samples=100):
"""
Compute counterfactual with uncertainty via bootstrapping.
"""
results = []
for _ in range(n_samples):
# Add noise to inference
cf_result = cf_engine.counterfactual(sample_idx, intervention)
results.append(cf_result['counterfactual']['symptom_severity'])
return {
'mean': np.mean(results),
'std': np.std(results),
'ci_lower': np.percentile(results, 2.5),
'ci_upper': np.percentile(results, 97.5)
}
3. Combine with Domain Knowledge
The most powerful counterfactuals come from:
Causal structure (DAG)
Domain expertise (mechanisms)
Data (observations)
Don't rely on any one alone.
You've Mastered Counterfactuals
Congratulations! You now understand:
✅ What counterfactuals are and why they're powerful
✅ The three-step process: Abduction → Action → Prediction
✅ How to implement counterfactual inference
✅ Individual Treatment Effects (ITE)
✅ Applications: personalization, explanation, regret analysis
✅ Counterfactual fairness
This is Level 3 reasoning. Most AI can't do this.
What's Next: Intervention Design
In Part 4 (Wednesday, Jan 22), we move from analysis to action:
How do we use counterfactuals to design optimal interventions?
What's the best treatment for each individual?
How do we optimize for multiple objectives?
How do we account for costs and constraints?
We'll build a complete intervention recommendation engine that combines everything from Parts 1-3.
Your Homework
1. Implement counterfactual engine
Use the code from this article
Test on your plant disease data
Generate counterfactual explanations
2. Experiment with interventions
Try different intervention values
Compare factual vs counterfactual outcomes
Find cases with high regret
3. Think about your domain
What counterfactual questions would be valuable?
What interventions do you want to optimize?
What constraints matter in practice?
4. Challenge yourself
Can you extend the engine to multiple treatments?
How would you handle continuous outcomes?
What about time-series counterfactuals?
Bring these to Part 4. We're building the intervention engine.
Series Navigation:
Part 3: Counterfactual Reasoning ← You are here
Part 4: Intervention Design → (Jan 22)
Part 5: Distributed Systems (Jan 24)
Code & Resources:
Part of the NeoForge Labs research series on production-grade causal AI.
Questions? I read every comment.